Supplemental Material for: A diffusion MRI tractography connectome of the mouse brain and comparison with neuronal tracer data
Evan Calabrese, Alexandra Badea, Gary Cofer, Yi Qi, G. Allan Johnson
Center for In Vivo Microscopy, Duke University Medical Center
Published in Cerebral Cortex (E-pub ahead of print) June 5, 2015 (PMID: 26048951) - Free full text through Open Access
Interest in structural brain connectivity has grown with the understanding that abnormal neural connections may play a role in neurologic and psychiatric diseases. Small-animal connectivity mapping techniques are particularly important for identifying aberrant connectivity in disease models. Diffusion MRI tractography can provide non-destructive, 3D, digital, brain-wide connectivity maps, but has historically been limited by low spatial resolution, low signal-to-noise ratio, and the difficulty in estimating multiple fiber orientations within a single image voxel. Small-animal diffusion tractography can be substantially improved through the combination of ex-vivo MRI with exogenous contrast agents, advanced diffusion acquisition and reconstruction techniques, and probabilistic fiber tracking. Here, we present a comprehensive, probabilistic tractography connectome of the mouse brain at microscopic resolution. This work serves as a reference database for future tractography studies in the mouse brain, and demonstrates an automated pipeline for assessing brain-wide connectivity in mouse models of human brain diseases.
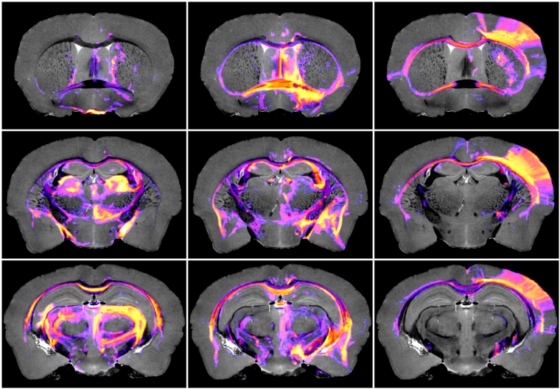
- View Tractography vs ABA Injectome Online in 3D
Access the CIVM online data visualization page, where you can interactively visualize and compare probabilistic tractography data and Allen Brain Atlas neuronal tracer connectivity data in 3D. Please watch the tutorial video below for help with controls. - View 2D Tractography Data Online in CIVMVoxPort
- Download Datasets in CIVMVoxPort
Download all of the data associated with this study including:
• Raw Image Data
• Estimated Fiber Data (BEDPOSTX)
• Probabilistic Tractography data (PROBTRACKX)
• Connectivity Matrix Graph Data
• Data Analysis/Processing Code
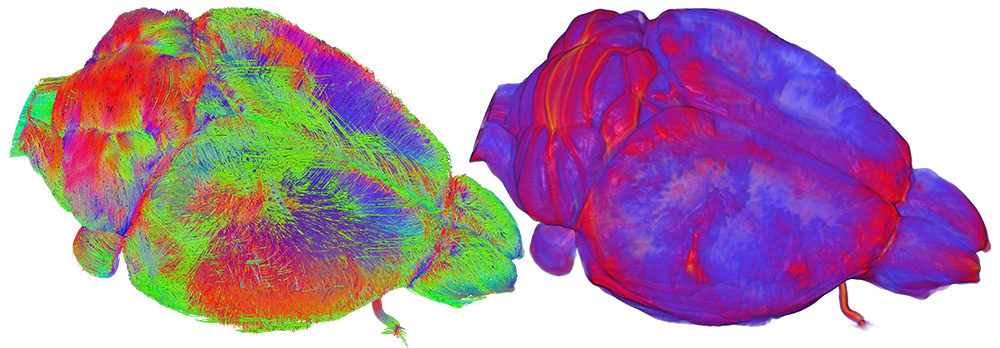
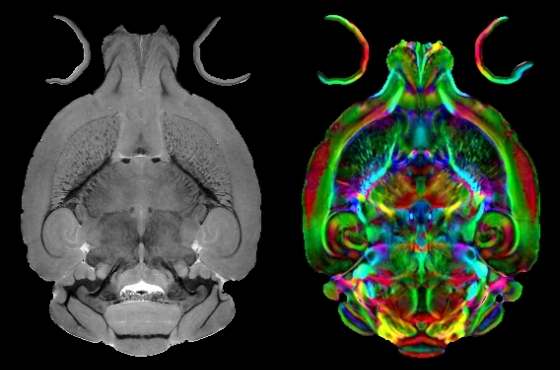
Use of CIVM Data:
Data downloaded from this site is for academic use only. If you use this data in a publication please send us a request for copyright permission and appropriate acknowledgements. Licenses can be granted for commercial use. Contact the Center for permission.
CIVM makes all data from published studies available for research. We ask that you provide contact information, and agree to give credit to the Duke Center for In Vivo Microscopy for any written or oral presentation using data from this site. Please use the following acknowledgement: Imaging data provided by the Duke Center for In Vivo Microscopy NIH/NIBIB (P41 EB015897) with additional support from (1S10OD010683-1).
Access Data in CIVM Voxport:
CIVM makes many types of data acquired for published and yet unpublished studies available through our CIVM VoxPort application. Use of VoxPort is free. Registration is required. Register for VoxPort access now. A new browser window or tab will open.
View Conventional MR imaging in CIVMVoxPort
System Requirements:
• CIVMVoxPort is designed to work on most platforms and is supported in most browsers.
• Registration with a valid email address is required to access Voxport.
• Login to Voxport is required to access available shared data sets.
• The ability to upload data or images into Voxport is currently disabled.
If you have trouble accessing CIVMVoxPort, contact us at CIVM Webmaster.
Acknowledgments:
All imaging was performed at the Duke Center for In Vivo Microscopy, an NIH/NIBIB National Biomedical Technology Resource Center (P41 EB015897). This project was also supported by NIH/Office of the Director (1S10OD010683-1).
